- Blog
- Art history volume 1 stokstad second edition
- Compiling java files on mac
- Cracked paint tool sai 64 bit windows 10
- Bioedit software online
- Samsung portable ssd t1 factory reset
- Just cause 2 mods ofw
- Miracast vs widi
- God hand game ign
- Trackmania 2 stadium just hold forward
- The revenant movie in hindi watch online
- Usbutil download english
- Moontide quartet kazim
- The office season 8 finale
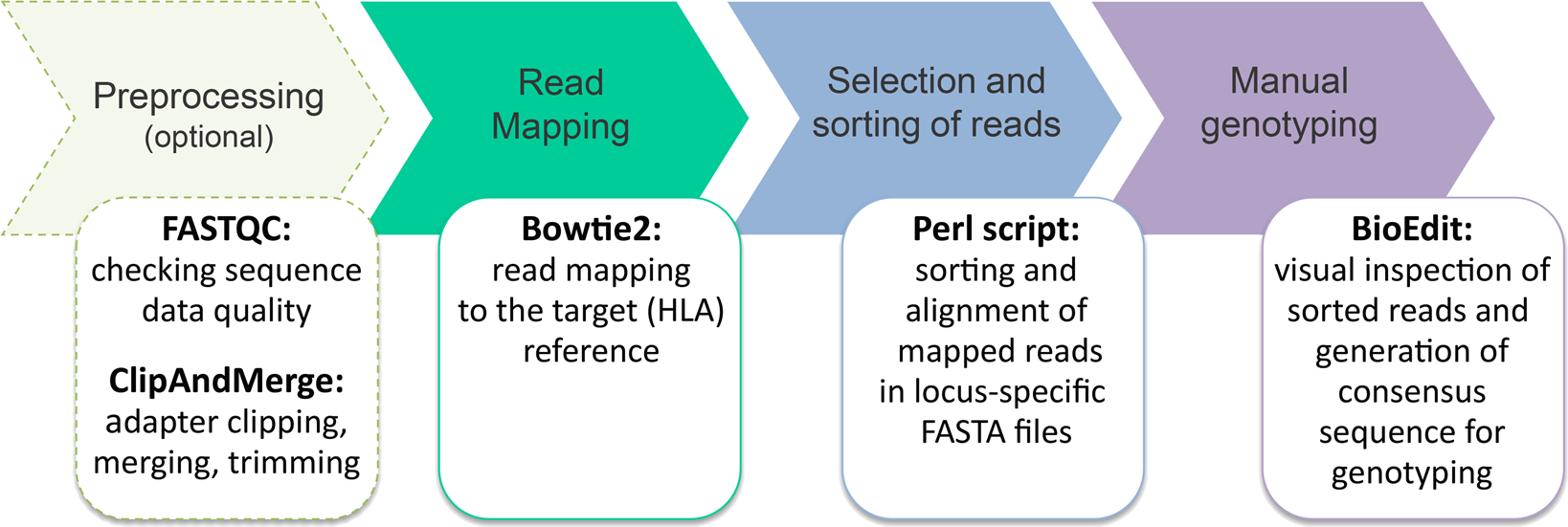
The web-site provides user-friendly web interface and user manual describing the underlying principle of the program.
The most widely used program for primer designing is PRIMER 3.0 ( ) with several versions of web interface. Orphan crops lack sequence information in which comparative genomics approaches such as homologous sequences are used to design degenerate primers, or re-sequence the gene of interest. Such cases include, but not limited to, retrieved sequences containing simple sequence repeats suitable for SSR marker development. There are several applications in which primer designing is required for marker development.
#Bioedit software online how to#
Information on how to access BLAST services on WWW, choosing the right type of BLAST, interpreting BLAST results, how to do batch BLAST jobs, and others can be found at NCBI-BLAST home page ( ). NCBI and EBI provide many different types of BLAST. BLAST is a set of programs used to compare a nucleotide or protein query sequence to all of the available sequence databases. The most common and widely used similarity search tool is BLAST (Best Local Alignment Search Tool (Ye et al. Sequence comparison is essential for understanding evolutionary relationship between genes.

In addition, web services such as EMBL-EBI provide similar tools. For beginners, stand alone programs such as MEGA can do excellent job of phylogeny tree construction. A deluge of tools and web services can also be found online (e.g. This is a broad topic and a subject of 100s of articles and books. In the context of this paper, phylogenetic analysis is performed to cluster multiple sequences based on genetic distances. Phylogenetic analysis is the basis of taxonomical and evolutionary studies. EMBL-EBI) using the ClustalW program or other methods. However, sequence alignment can also be done on the web at one of the resources listed in this review (e.g. A pair of sequences or multiple sequences saved, for example in FASTA format, can be used as an input. 2007), are freely downloaded and installed with easy-to-understand user’s manual. Two examples of widely used open access softwares, namely BioEdit (Hall, 1999), and MEGA (Tamura et al. Various algorithms have been developed to produce optimal alignment, a topic which is beyond the scope of this paper. Sequence alignment is the prerequisite of virtually all forms of sequence analysis ranging from search, to assembly, and to phylogenetics. The result can be displayed in different format or downloaded, the most common format being FASTA. Search can be performed in all databases or restricted to nucleotide in the drop-down menu. The Entrez search engine at NCBI, in addition to retrieving sequences, returns pre-computed lists of data elements such as related sequences, gene, protein, taxonomy, and others. Search for a sequence of interest begins with keywords, accession number, gene name, species name, etc.
